Gene expression analysis by sequencing
Acobiom provides RNA-Seq analysis services. This approach allows to measure the expression level of genes by sequencing, an expertise of over 20 years of the company.
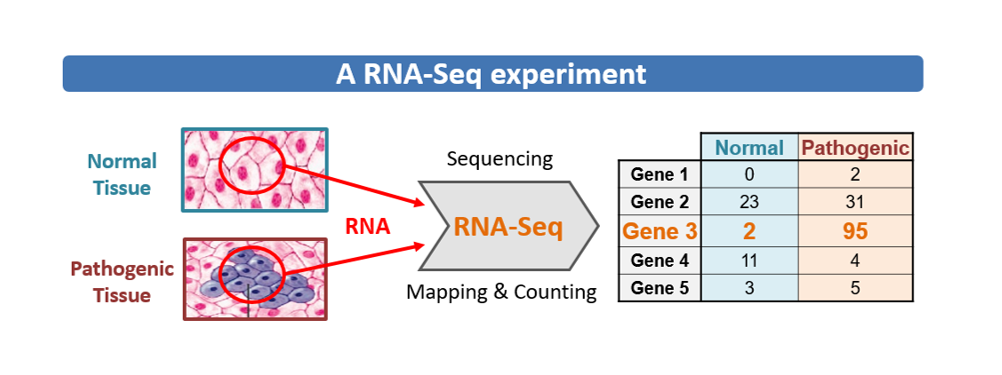
RNA-Seq is a powerful method to identify and quantitatively decode the entire population of RNAs in a sample.
This gene expression (transcriptome) analysis allows to determine which genes are expressed under different conditions and to compare their expression levels.
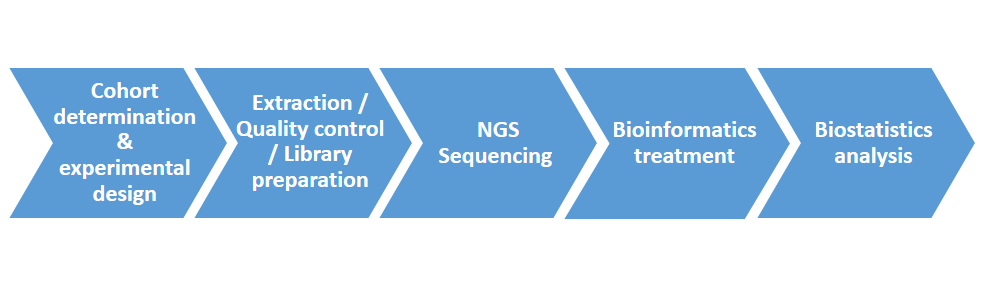
From study design advice, through RNA extraction, bioinformatics processing of the generated sequences, and biostatistical analysis of the obtained transcriptomic data, Acobiom provides a complete pipeline described in the workflow below.
RNA-Seq is used in a wide variety of applications: identification of
biomarkers linked to a pathology or a physiopathological condition, analysis of
the effects of a molecule or a treatment, stratification of patients to target
a therapy…
Moreover, the RNA-Seq technique does not necessarily depend on prior knowledge of the sequences or the genome. Therefore, transcriptome profiling of any species is possible.
An over 20-year expertise dedicated to each study
The over 20-year expertise of Acobiom in performing transcriptome by sequencing enables it to produce high-quality, low-cost libraries, especially when working with small amounts of total RNA (50 to 100 ng).
The NGS sequencing step is carried out via an Illumina sequencer the most adapted to the project, in order to optimize the results of each study according to the number of samples, the desired reading depth…
The company then proposes the use of specific bioinformatics tools and personalized algorithms according to the project and the expectations of its customers and partners.
For each RNA-Seq analysis, Acobiom proposes to perform the steps
described below or some of them, depending on the project:
- Experimental design,
- Optimization of sampling and storage preparation,
- RNA extraction and qualification,
- Library preparation,
- Sequencing,
- Data Quality check,
- Mapping on reference genome,
- Comparison of read counts,
- Annotation of sequences,
- Overview of expressed genes including statistics analysis, visualization and validation.
Bioinformatic processing of RNA-Seq data
For the bioinformatic analysis of the transcriptome by RNA-Seq, Acobiom follows a workflow including the following steps:
- Gene expression profiling for different biological conditions (mapping on an annotated reference genome, counting, normalization, sequence annotation),
- Overview and comparisons of expressed genes (differential analyses, visualization and statistical validation) .
- Control the quality of the sequencing data;
- Clean the sequence data;
- Map these data on a reference genome;
- Count the mapped sequences;
- Standardize the data;
- Compare the data with other RNA-Seq profiles;
- Annotate and analyze the data;
- Visualize and validate statistically the results.
Deliverables include:
- Raw sequences (FASTQ),
- Mapping results (BAM),
- Variant tables (VCF), annotations,
- Statistical analysis to identify variants of interest.
The RNA-Seq provides absolute values and does not require calibration with arbitrary standards. The transcriptomic profiles obtained can be compared to other RNA-Seq profiles generated by the scientific community or other laboratories, and in particular those present in the MaRS database developed by the company.
For more information on the RNA-Seq services provided by Acobiom, please contact the company.

